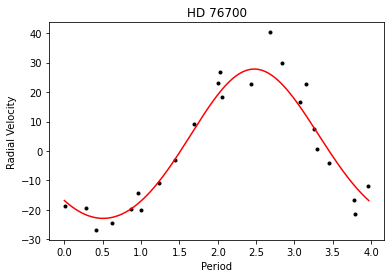
HD 76700
Radial velocity data for HD76700 were obtained from Tinney et al.(2003).Author: Do Kester
We need to import some classes
In[1]:
import numpy as numpy import math from astropy.io import ascii from BayesicFitting import RadialVelocityModel from BayesicFitting import PolynomialModel from BayesicFitting import CircularUniformPrior from BayesicFitting import NestedSampler from BayesicFitting import formatter as fmt from BayesicFitting import plotFit from BayesicFitting import Tools import matplotlib.pyplot as plt
In[2]:
#%matplotlib osx
Read the data
In[3]:
tbl = ascii.read( "data/HD76700-Tinney.dat" ) #print( tbl ) jd = tbl['JDa'].data rv = tbl['RVa'].data er = tbl['Uncertainty'].data wgt = 100.0 / numpy.square( er )
We define a radial velocity model. It has 5 parameters: eccentricity,
amplitude, period, phase of periastron, longitude of periastron. The
phase parameters both get a CircularUniformPrior. We need to add a
constant for the systemic velocity of the system.
In[4]:
twopi = 2 * math.pi rvm = RadialVelocityModel( ) lolim = [0.0, 0.0, 3.0] hilim = [0.9, 40.0, 5.0] rvm.setLimits( lowLimits=lolim, highLimits=hilim ) rvm.setPrior( 3, prior=CircularUniformPrior(), limits=[0,twopi] ) rvm.setPrior( 4, prior=CircularUniformPrior(), limits=[0,twopi] ) #Tools.printclass( rvm ) pm = PolynomialModel( 0 ) pm.setLimits( lowLimits=[-20], highLimits=[20] ) #sm *= hm mdl = pm + rvm print( mdl )
Out[4]:
Polynomial: f( x:p ) = p_0 + RadialVelocity
In[5]:
# define NestedSampler ns = NestedSampler( jd, mdl, rv, weights=wgt, seed=1301 ) ns.ensemble = 500 # set limits on the noise scale of the distribution ns.distribution.setLimits( [0.01,100] ) # run NestedSampler evi = ns.sample( plot=True )
Out[5]:
Fit all parameters of Polynomial: f( x:p ) = p_0 + RadialVelocity Using a Gauss error distribution with with unknown scale Moving the walkers with GalileanEngine ChordEngine >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> >>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>>> >>>>>>>>>>>>>>>>>>>>>>>>>>>>>> Iteration logZ H LowL npar parameters 27978 -353 28.0 -321 7 [ 0.724 0.075 25.647 3.971 3.805 6.159 4.128] Engines success reject failed best calls GalileanEngine 159283 56852 96987 58 30362 ChordEngine 212502 1126682 2 43 30362 Calls to LogL 1652308 to dLogL 56852 Samples 28478 Evidence -153.510 +- 0.103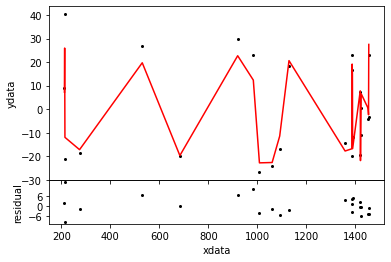
In[6]:
sl = ns.samples par = sl.parameters std = sl.stdevs print( fmt( par, max=None ) ) print( fmt( std, max=None ) ) pal = par.copy() stl = std.copy() pal[5] *= 180 / math.pi pal[4] *= 0.5 * pal[3] / math.pi stl[5] *= 180 / math.pi stl[4] *= 0.5 * pal[3] / math.pi print( fmt( pal, max=None ) ) print( fmt( stl, max=None ) ) print( fmt( sl.scale ), fmt( sl.stdevScale ) ) print( evi )
Out[6]:
[ 0.758 0.067 25.342 3.971 3.854 6.207]
[ 0.528 0.026 0.755 0.000 0.492 0.493]
[ 0.758 0.067 25.342 3.971 2.436 355.662]
[ 0.528 0.026 0.755 0.000 0.311 28.219]
4.268 0.339
-153.51027291941526
In[7]:
plt.plot( jd % par[3], rv, 'k. ' ) xx = numpy.linspace( 0, par[3], 1001, dtype=float ) plt.plot( xx, mdl.result( xx, par ), 'r-' ) plt.xlabel( "Period") plt.ylabel( "Radial Velocity") plt.title( "HD 76700" ) plt.show()
Out[7]:
In[9]:
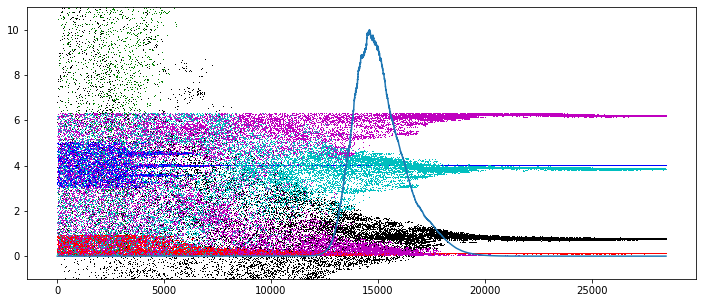
plt.figure( 2, figsize=[12,5] )
pev = sl.getParameterEvolution()
c = ['k,', 'r,', 'g,', 'b,', 'c,', 'm,']
for k in range( mdl.npars ):
plt.plot( pev[:,k], c[k] )
wev = sl.getWeightEvolution()
wev /= numpy.max( wev )
plt.plot( wev * 10 )
plt.ylim( -1, 11 )
plt.show()